BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:
URL: http://ijmm.ir/article-1-1590-en.html

 , Mohsen Moghoofei2
, Mohsen Moghoofei2 
 , Ahmad Tavakoli1
, Ahmad Tavakoli1 
 , Alireza Tabibzadeh1
, Alireza Tabibzadeh1 
 , Saied Ghorbani1
, Saied Ghorbani1 
 , Hadi Ghaffari3
, Hadi Ghaffari3 
 , Shayan Mostafaei4
, Shayan Mostafaei4 
 , Seyed Hamidreza Monavari5
, Seyed Hamidreza Monavari5 

2- Department of Microbiology, Faculty of Medicine, Kermanshah University of Medical Sciences, Kermanshah, Iran
3- Department of Bacteriology and Virology, Semnan University of Medical Sciences, Semnan, Iran
4- Department of Biostatistics, School of Health, Kermanshah University of Medical Sciences, Kermanshah, Iran
5- Department of Medical Virology, School of Medicine, Iran University of Medical Sciences, Tehran, Iran ,
The human immunodeficiency virus (HIV) infects 37.9 million people worldwide and is a significant global health concern (1, 2). HIV infection reduces T lymphocyte CD4+ cells and progresses toward immunodeficiency (3). Co-infection of hepatitis B virus (HBV) with HIV is a significant health problem due to the acquisition risk factors (4). The infection rate of HBV in HIV-positive patients is variable and depends on high-risk groups, socioeconomic and geographical regions (5, 6). It has been estimated that 2.7 million people worldwide are co-infected with HIV and HBV (7). Different diseases or infections could affect the clinical course of HBV infection (8-11). For instance, HIV accelerates the progression of HBV infection (12, 13). However, the underlying mechanisms of the accelerated progression to liver cirrhosis due to HIV/HBV co-infections are not fully understood (14). The mortality rate of liver diseases in HIV/HBV co-infected patients is 17 times greater than in HBV mono-infection (14). Meanwhile, the impact of HBV on the natural history of HIV infection is still unknown (15).
MicroRNAs are non-coding RNA with a length of 18-24 nucleotides and can induce post-transcription suppression or reduce the expression of target genes (16-18). Recently, the discovery of miRNAs as novel biomarkers in serum or plasma has represented a new approach for the diagnostic area (18). The reports show that microRNAs can regulate replication, latency, pathogenesis, and increased immune responses against viral infections (19).
It has been reported that miR-122 accounts for about 50%-70% of the total liver miRNAs. This particular miRNA is critical in lipid metabolism (20). Also, miR-122 could reduce the replication rate of HBV through interaction with the HBV genome (21). Furthermore, other miRNAs can interfere with viral pathogenesis or replication stages. For instance, miR-149 can block the Vpr protein of HIV through direct interaction (22, 23). The current study aimed to investigate the expression level of the miRNA-122, miRNA-149, miRNA-199a, and let7 in HIV, HBV, HIV/HBV co-infection patients, and healthy controls.
Study Population
The present study has a cross-sectional design. Due to the low frequency of HBV-HIV co-infected patients and the increased likelihood of identifying this co-infection, about 200 HIV-infected patients were first enrolled in this study. The following formula was then used to calculate the sample size:
Four study groups were in this investigation, including 13 HIV-positive patients, 13 HBV-positive patients, 13 patients with HIV-HBV infection, and 13 healthy individuals. The study was approved by the ethics committee of the Iran University of Medical Science (Ethical Code: IR.IUMS.FMD.RC.1398.169), and informed consent was obtained from each participant.
The subjects included in the present study were patients with a new diagnosis of HBV and HIV infections who did not receive therapy. In fact, they were identified as a new case. Also, subjects in all age groups were included in the present study. Another criterion for inclusion in the study was the absence of other infections. Patients treated with different drugs against HBV and HIV were excluded from the study.
Peripheral blood mononuclear cells (PBMC) samples were collected from all four groups. The patient's history was investigated for other background or acute diseases like HCV, mycobacterium tuberculosis, or cytomegalovirus (CMV) infection. We have tried to match the groups for sex, age, and other demographical variables.
Serologic Assessment
The HBV was confirmed by assessing the HBsAg in the serum sample of the enrolled patients. Based on the manufacturer's protocol, the HBsAg was evaluated by a commercial ELISA kit (DIA.PRO, Milano, Italy). The fourth-generation commercial ELISA kit (DIA.PRO, Milano, Italy) was also used to confirm HIV infection.
PBMC Isolation
PBMC samples were isolated from all samples by Ficoll gradient separation using FicollHypaque (Lympholyte-H, Cedarlane, Hornby, Canada) density gradient centrifugation. The PBMCs were washed three times with phosphate-buffered saline (pH 7.3 ± 0.1) and stored at -20ºC (23).
DNA and RNA Extraction
To assess HBV and HIV viral load, DNA and RNA were extracted by the GeneProof PathogenFree DNA nucleic acid extraction kit (GeneProof, Brno, Czech Republic) and Favorgen RNA extraction kit (Favorgen Biotech), respectively. Also, according to the manufacturer's instructions, total RNA was isolated from PBMCs using RNX™- Plus Kit (Sinagene, Iran). First, the BON-miR miRNA 1st-strand cDNA synthesis kit (Bonyakhteh, Tehran, Iran, Cat No # BON209001) was used for the polyadenylation of microRNAs and then, the polyadenylated microRNAs were converted into cDNA.
Viral load and T-lymphocyte CD4+ Count
The HIV viral load was determined using the QIAamp DSP Virus Kit (Qiagen GmbH, Hilden, Germany), based on the manufacturer's protocols. According to the manufacturer's instructions, HBV viral load was also measured using the GeneProof Hepatitis B Virus (HBV) PCR kit (GeneProof). CD4+ count was determined by the PIMA CD4+ analyzer (Alere, Germany).
Real-time RT-qPCR
The real-time RT-qPCR assay was used to quantify the level of the miRNAs, and SNORD68 was used as an internal control and relative quantification. All reactions were performed in duplicate.
Of cDNA 1 µg, 2X Master Mix (QIAGEN GmbH, Hilden, Germany), 1 µM of each primer (Metabion, Munich, Germany), and nuclease-free water up to 20 µl made up the reaction mixture. The following were the ideal real-time PCR conditions: Denaturation step at 95°C for 2 minutes, followed by 40 cycles of 95°C for 5 seconds and 60°C for 30 seconds. Table 1 lists the primers that were used in this study.
Statistical Analysis
In this study, the expression level and quantification for each microRNA were analyzed using the 2-ΔΔCT. Data were analyzed by χ2 or Fisher's exact test using SPSS software Version 22.0 (IBM Corp. Released 2013. IBM SPSS Statistics for Windows). Correlations between serum miRNA-122, miRNA-149, miRNA-199a, miRNA- let7, and viral load were analyzed by Spearman's correlation.
Table 1. Primers for miRNAs amplification
| miR | Forward primer | Reverse primer |
| miR-149 | 5'-GGCTCTGGCTCCGTGTCTT-3' | 5'-CAGTGCAGGGT CCGAGGTATT-3' |
| miR-199 | 5'-GCATAGCCCGCCCAGTGTT-3' | 5'-GTGCAGGGTCCGAGGT-3' |
| miR-let7 | 5'-GTCTGGTCAGAATC ACCTC-3' | 5'TTCTGCGTGTCTCC AACTCA-3' |
| miR-122 | 5'- GCTCGACCTCTCTA TGGGC-3' | 5'-TTAAGCCCTGCGTG TCTCC-3' |
| SNORD68 | 5’-CAAATTCGTGAAGCGTTCCATA-3’ | 5’-AGTGCAGGGTCCGAGGTA TTC-3’ |
Table 2. Participants' demographic characteristics; patients, were in three groups, including HIV, HBV, and HIV-HBV co-infected, and a control group was considered.
| Characteristics | Control (N=13) | HIV (N=13) | HBV (N=13) | HIV & HBV (N=13) | P- value |
|
| Age (Year)* | 33.75 ± 7.35 | 35.70 ± 8.20 | 37.80 ± 11.04 | 33.50 ± 10.13 | 0.433 | |
| Sex+ | Male | 7 (53.8%) | 10 (76.9%) | 8 (61.5%) | 9 (69.21%) | 0.456 |
| Female | - | - | - | |||
| CD4 (count)* | 1584.5 ± 631.87 |
454.8 ± 304.04 | 868.9 ± 209.22 | 265.6 ± 151.11 | <0.001 | |
| HIV load (count)* | NA | 3985291.7± 2385361.18 |
NA | 1400362.8± 1226925.03 |
0.341 | |
| HBV load (count)* | NA | NA | 10401116.2± 5152977.67 |
6134541.0± 3717689.46 |
0.506 | |
Table 3. Comparison of microRNAs expression levels between HIV, HBV, and HIV-HBV co-infected groups compared with the healthy control group.
| Expression Level | Control | HIV | HBV | HIV & HBV | F, adj.P | F, adj.P | F, adj.P |
| miR-149 | 1.89±0.826 | 1.78±0.81 | 4.27±2.42 | 4.14±1.97 | 0.94, 0.997 | 2.26, <0.001 | 2.19, <0.001 |
| miR-199 | 6.18±2.59 | 3.68±2.50 | 2.09±0.89 | 2.11±0.80 | 0.59, <0.001 | 0.34, <0.001 | 0.34, <0.001 |
| miR-let7 | 7.95±3.01 | 1.81±0.65 | 2.10±0.98 | 1.72±0.47 | 0.23, <0.001 | 0.26, <0.001 | 0.22, <0.001 |
| miR-122 | 1.81±0.63 | 2.43±1.10 | 3.51±1.53 | 5.55±2.95 | 1.34, 0.692 | 1.94, 0.018 | 3.07, <0.001 |
Table 4. Spearman's correlation coefficient between microRNAs expression level with HIV loads, age, and CD4 count in HIV+ patients.
| Age | HIV load |
CD4 count |
|
| miR-149 | 0.209 | -0.58*** | -0.002 |
| miR-199 | -0.274 | 0.131 | 0.001 |
| miR-let7 | -0.044 | -0.274* | -0.294* |
| miR-122 | -0.335 | -0.203* | 0.248* |
Table 5. Spearman's correlation coefficient between microRNAs expression levels with HBV and HIV loads, age, and CD4 count in the patients.
| Age | HIV load |
HBV load |
CD4 count |
|
| miR-149 | 0.061 | -0.27* | -0.121 | 0.044 |
| miR-199 | -0.11 | 0.058 | 0.427** | -0.127 |
| miR-let7 | -0.107 | -0.228* | 0.051 | 0.015 |
| miR-122 | -0.134 | -0.064 | 0.067 | -0.136 |
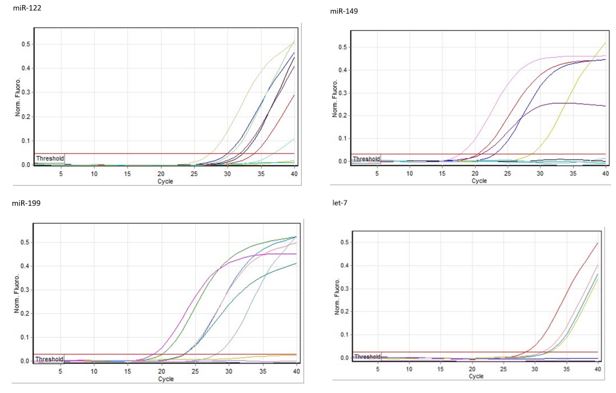
Figure 1. Real-time PCR curves for microRNAs expression levels between HIV, HBV, and HIV & HBV co-infected groups with healthy controls.
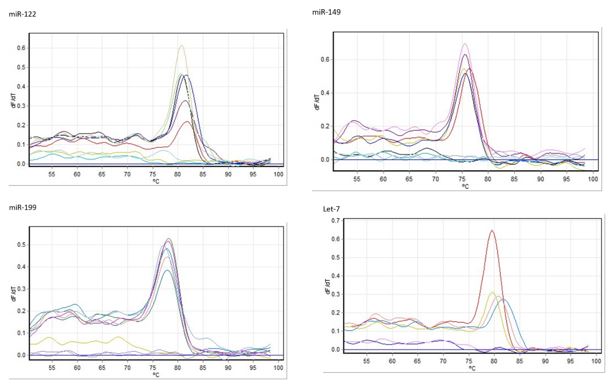
Figure 2. The melting curves for each interested microRNA.

Figure 3. Schematic diagram of comparison between expression levels of each interested microRNA among HIV, HBV, and HIV-HBV co-infected patients.
The miRNAs are important elements in the post-transcriptional regulation of gene expression (24), and they have been recently introduced as an important element in host-pathogen interactions (25). It has also been suggested that the miRNAs can serve as potential biomarkers for diagnosis and as a prognosis factor (26). MiR-let7 is a tumor-suppressor miRNA that targets well-known oncogenes like Ras, STAT3, and Myc. MiR-122 is one of the most common miRNAs in the liver, accounting for over 70% and 52% of the entire hepatic miRNome in adult mice and humans, respectively. The microRNA miR-122 is important for liver development, differentiation, homeostasis, and function. In the adult liver, MiR-122 is also important for cholesterol and fatty acid metabolism control (27). Furthermore, miR-122 has been demonstrated to indirectly interfere with HBV replication by lowering cyclin G1 expression, leading to p53-mediated HBV transcription suppression. However, miR-122 was found to indirectly promote HBV replication in human hepatoma cell lines by suppressing HMOX1, which interfered with HBV replication by lowering the stability of the HBV core protein (28). Through binding to the HBsAg-coding region, MiR-199a-3p can effectively limit viral replication by lowering HBsAg expression (29). In the current study, we evaluated the expression level of miRNA-122, miRNA-149, miRNA-199a, and miRNA-let7 in PBMCs of HBV, HIV, and HIV/HBV co-infected patients and healthy controls.
Previous studies suggested that HBV x protein can induce a faster progression to the AIDS phase in HIV-positive patients and reduce the survival rate in HIV/HBV Co-infections patients (16-18, 30). Investigations indicated that the HBV/HIV co-infections increase the HBV infectivity and risk of cirrhosis progression (14, 31).
MiR-122 plays a critical role in inhibiting HBV replication (32). This miRNA induces HBV inhibition by direct binding into the HBV polymerase region (33). Ebrahimifard et al. (34) demonstrated that the miR-122 is upregulated in patients with chronic hepatitis B compared to the control group. Also, Ji et al. (35) indicated that the expression of the liver-specific miR-122 was significantly upregulated in HBV-infected patients. In contrast, Wang and colleagues (27) showed that the expression level of miR-122 was lower in HBV infection than in healthy controls. Furthermore, early investigations have shown that the deregulation of this miRNA is linked to the progress of HCC (36). Yousefpouran and colleagues (37) assessed the miRNAs expression in HIV, HCV, HBV, HIV-HCV, and HIV- HBV co-infection patients. Yousefpouran's study indicated that miRNA 29, 34, 122, 128, 146, 155, and 181 were upregulated in HIV-HBV co-infected patients compared with healthy controls, while miRNA 22 and 223 were down-regulated in HIV-HBV co-infected patients in comparison with healthy controls. Considering these results, it seems that the current study results are consistent with the results of conducted studies in this field. It should be noted that a major limitation in our current study was the limited sample size. Also, further studies in this field are required for a clear conclusion.
The current study showed that the expression level of miR-149 in patients with HBV/HIV co-infected and HBV-infected patients was increased. It has been demonstrated that miR-149 can induce apoptosis in THP-1 cells (Tohoku Hospital Pediatrics-1) (38). Also, miR-149 can target HIV Vpr protein (39). The miRNA-149 is an oncogene regulator miRNA, but its exact role in HCC or HBV infection is unclear. But the polymorphisms in this miRNA seem to be associated with the HBV-associated HCC progression (40, 41). The study by Squillace et al. (42) also indicated that the miRNA-149 was upregulated in HIV patients in subcutaneous adipose tissue compared to healthy controls. The current report results seem to be in line with the mentioned studies. But further studies in this manner would be necessary.
Our study also showed that the expression level of mir-199 and mir-let7 decreased in all patients compared to the healthy subjects (P<0.001). It has been reported that downregulation in the expression of miR-let7 is associated with a reduction in HBV replication and HCC progression (43, 44). These two miRNAs are essential regulators of development, differentiation, and cell proliferation (45). Similar to our results, Wang et al. (46) demonstrated miRNA-let7 downregulation in cell lines and HBV-infected patients. This down-regulation results in suppressed cell proliferation and potentially viral replication and hepatocarcinogenesis (46). Swaminathan et al. (47) demonstrated that let-7 miRNAs decreased following HIV-1 infection in HUT 78 cells in another study. Most of the mentioned studies support our findings of the miRNA-let7.
The miR-199 could be affected due to the HBV infection. The miR-199 is an important miRNA in triggering antiviral signaling pathways (48, 49). The exact role of miR-199a, like the mir- 149, in liver diseases and HIV infection remains controversial (21). Hou et al. (50) demonstrated that miR-199a/b-3p is consistently decreased in HCC. In contrast, Zhang et al. (48) identified an increase in miR-200 and miR-199 in HepG2 2.2.15 cells. Also, Zhang et al. suggested that this miRNA can suppress HBsAg expression and HBV proliferation. Studies show that some miRNAs, such as miR-122 and miR-199 family members, have high expression in the liver and exhibit HBV-mediated expression changes (51). So the change in the expression of these microRNAs can have an important effect on the host cell and virus (19).
The mentioned studies in this field mostly confirmed our results. There are slight differences between the current study result and the mentioned studies. These differences could be justified based on the sample size and methodology differences. The major limitation of this study was due to the sample size, which can be suggested for further investigation to design more progressive studies in this field. The conducted study by Huang et al. (52) systematically reviewed and Meta‐ Analysis the potential circulating miRNAs in HCC diagnosis. The results suggested miR-21 and miR-122 as promising biomarkers due to the great sensitivity and specificity of these miRNAs. MiRNAs could be a great tool for disease progression and diagnosis (53). But it has been suggested that the miRNAs with an abundant high-level of expression could be a target as therapeutic agents (54). Furthermore, further miRNAs therapeutic potential is not neglectable. For instance, Weidhaas and colleagues (1) indicated the great potential in using miRNAs as tools for the suppression of the resistance to the anti- cancer cytotoxic radiation therapy.
In summary, due to the relative stability of microRNAs in cells and body fluids, different expression levels at different stages of the disease, and even being specific to a particular disease, these genetic factors can be used as markers to diagnose, monitoring of disease progression, and even monitoring of response to therapy. The present study is the first report to determine the expression levels of miRNA-122, miRNA-149, miRNA-199a, and mir-let7 in PBMC and serum samples of patients co-infected with HIV/HBV infection in Iran. The expression levels of microRNAs in the pathogenesis of HIV/HBV co-infection at different stages of the disease and the manipulation of these microRNAs can indicate a new pathway for controlling HIV/HBV co-infection, which requires more investigations with a larger sample size.
We would like to acknowledge the staff of the virology department of Iran University of Medical Sciences for their cooperation.
All procedures performed in this work followed the ethical standard of the committee of the Iran University of Medical Sciences (Ethical Code: IR.IUMS.FMD.RC.1398.169), and informed consent was obtained from each participant.
SH.M designed and administrated the project. SSM and Ah.T wrote the manuscript. SSM, M.M and Al.T performed the experiment. S.M and MM analyzed the data. S.G and H.G collected patient’s data and samples. All authors read and approved the final version of the manuscript.
This project was financially supported by the Iran University of Medical Sciences (Grant number: 13-359).
Conflicts of Interest
The authors have no conflict of interest.
Received: 2021/12/20 | Accepted: 2022/05/21 | ePublished: 2022/08/8
| Rights and permissions | |
 |
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License. |






