BibTeX | RIS | EndNote | Medlars | ProCite | Reference Manager | RefWorks
Send citation to:
URL: http://ijmm.ir/article-1-1324-en.html
2- Department of Biology, Faculty of Basic Sciences, University of Maragheh, Maragheh, Iran
Pseudomonas aeruginosa is a gram-negative, mobile and opportunistic pathogenic bacillus. Due to its high compatibility and inherent resistance to various antibiotics, this microorganism is durable in different environmental conditions, such as different levels in medical centers. P. aeruginosa is a major cause of nosocomial infections, and the prevalence of such infections is evident in cases of host immune system disorders, severe burns, or cystic fibrosis (1). Treatment methods are increasingly limited due to the continued emergence and development of strains resistant to antimicrobial agents. P. aeruginosa is one of the most common nosocomial pathogens and the second most common pathogen isolated from ventilator-associated pneumonia. P. aeruginosa infections have high morbidity and mortality (2). It is shown that the improper use of antimicrobials has caused the bacterium to be highly resistant to carbapenems (3).
Lytic bacteriophages with the ability to multiply in a bacterial culture can quickly remove bacterial infections, which are not easily treated with antibiotics, regardless of antibiotic resistance. Hence, lytic phages are ideal candidates for use as bio-therapeutic agents (4). In recent years, due to the increasing antibiotic resistance of new bacteria and the potential of phages to remove bacteria with multiple antibiotic resistance, new phages can be isolated, described, and screened against multidrug-resistant bacteria (5).
Phages are generally self-specific in attacking their target and can inhibit specific species or strains of bacteria. Nowadays, therapeutic and antibacterial applications of phages, especially in clinical, medical and veterinary fields, are known as phage therapy, to which attention has been paid (6). To evaluate the potential of isolated phages so as to fight bacteria and determine the optimal conditions of phage function for their use in the phage therapy process, a more detailed study of the characteristics of each bacteriophage is necessary. The aim of this study was to isolate and evaluate the efficacy and biological and morphological characteristics of specific phages against P. aeruginosa strains. Furthermore, this study evaluates their antibacterial potential against some infections caused by gram-positive and gram-negative bacteria.
Bacterial Strains
In this study, the standard strains of P. aeruginosa ATCC 27853, P. aeruginosa ATCC 9027, Klebsiella pneumoniae PTCC 1290, Staphylococcus aureus ATCC 25923, Salmonella enterica ATCC 19430 and Escherichia coli ATCC 8739, were used, all of which were purchased from Persian Type Culture Collection. In standard laboratory tests, P. aeruginosa ATCC 27853 and P. aeruginosa ATCC 9027 reference strains were used for quality control and antibiotic susceptibility testing. Also, to study the host range of isolated bacteriophages, clinical isolates of P. aeruginosa, K. pneumoniae, S. aureus, and E. coli were used, and all of them were isolated from different medical centers in East Azerbaijan of Iran from 2016 to 2018. Isolates were identified by using standard biochemical tests (7).
Isolation, Purification, and Titration of Phages
To isolate lytic bacteriophages against P. aeruginosa ATCC 27853 and ATCC 9027, several water samples were taken from Sufi Chai River and municipal sewage of Maragheh city (Iran) from May to June 2017. The samples (50 mL) were centrifuged at 1000 rpm for 5 min. The supernatant was passed through a filter (Jet Biofil, Canada) with a pore size of 0.45 μm. Then, 5 mL of LB medium (Himedia, India) with double concentration, 5 mL of 4 h bacterial culture, and 70 μL of MgSo4 (1% w/v) (Merck, Germany) (for optimal binding of phage to bacteria) were added to it. It was stored for 24 hours at 37°C in a shaker incubator (Vision, Korea) at 180 rpm. Then 3 mL of chloroform (Merck, Germany) was added to the solution and incubated at room temperature for 2 h.
The above solution was passed through a filter after centrifugation (30 min at 3500 rpm) (8). The formation of phage plaques was investigated by double-layer agar culture. In order to purification, a clear lytic plaque was removed from the medium by using a sterile Pasteur pipette and mixed with the LB culture medium containing the host bacteria, and incubated overnight at 37°C. For removing bacteria, chloroform was added and centrifuged. Then double-layer agar was prepared, and the plaques were examined in terms of shape and size. This step was repeated three times to obtain pure phages (9). Phage titration was performed according to the double-layer agar method. In summary, after dilution of phage solution, 100 μL of diluted phage solution, 250 μL of host bacteria, and 3 mL of molten LB containing 0.7% agar were mixed. It was poured on a plate comprising LB medium containing 1.5% agar. The plates were incubated overnight at 37°C. The number of phages in the medium was expressed as PFU/mL (10).
PFU/mL= Volume used × Reverse dilution used × Number of plaques.
Testing of the Optimal Multiplicity of Infection (MOI)
Overnight cultures of each P. aeruginosa strain were diluted 1:100 in fresh liquid LB and kept in an incubator at 37°C until reaching the primary logarithmic growth phase (OD600= 0.4-0.6). Then, the solution was mixed with 104 to 108 PFU/mL phage dilutions and incubated at 37°C for 3.5 h. The combination was centrifuged at 5000 g for 20 min at 4°C, and the supernatant was filtered through a 0.45 μm filter, and immediately double-layer agar was prepared (11).
Absorption Rate of Bacteriophage
Bacteria were grown in LB at 37°C until reaching OD600= 0.3 nm. 6 mL of each sample was centrifuged, and the pellets were washed with 1 mL of 0.85% NaCl (Merck, Germany). After re-centrifugation, each pellet was suspended in 1.3 mL of LB medium and incubated at 37°C for 15 minutes. Then, bacteriophage with MOI= 0.1 was added to it. The supernatant titer was determined in intervals during incubation using a centrifuging step for 1 min at 6000 g at room temperature. The plates were incubated overnight at 37°C, and the phage was added to the bacterial host (time zero) suspension immediately; samples were taken and considered as non-absorbed phages (100%) (12).
One-Step Growth Curve
The latent time and burst size of the bacteriophages were determined by a one-step growth curve. For this purpose, bacterial hosts were grown in LB medium at 37°C until reaching OD600= 0.2 nm. Then, 10 mL of the samples were centrifuged with 4000 g at 4°C for 10 min. The supernatant was removed, and each pellet was suspended in 1 mL of fresh LB medium with 3 mM NaN3 (Merck, Germany) and incubated at 37°C for 5 min. Then, the bacteriophage with an optimal multiplicity of infection (MOI) of 0.1 was added to the suspension. After 10 min of incubation, the non-adsorbed phages were removed by washing three times with an LB medium containing 3 mM NaN3. In the next step, 25 μL of phage suspension was added to 25 mL of LB medium and aerated at 37°C (time 0). Then, the number of phage-infected bacterial cells was determined 1 min after infection by mixing 5 μL of the sample with 0.995 mL of the overnight host culture using the double-layer agar method. Samples were collected every 1 min for the first 10 min and then every 2.5 min. Finally, successive dilutions were prepared from them, and the samples' titers were determined (12).
Morphology of Plaques and Phages
Plaques were examined for diameter, edges, and turbidity (13). In order to study the morphology of the isolated phages, each pure filtered phage (108 PFU/ mL) was negatively stained with (2% w/ v) uranyl acetate and fixed on a copper grade. To observe the bacteriophage morphology, a transmission electron microscope (Leo 906, Germany) operated at 100kV was used (14). Morphological features were used to classify both phages according to the guidelines of the International Committee on Taxonomy of Viruses (ICTV).
Survival of Bacteriophages at Different Temperature Conditions
To evaluate the stability of phages at different temperatures, 100 μL of phage solution was suspended (108 PFU/ mL) in 900 μL SM buffer, and the suspension was stored at -20, 4, 37, 22, 40, and 60°C. Samples were taken at 60 min and 24 h. The survival rate of each bacteriophage at any temperature was determined by the double-layer agar plating method (15).
Survival of Bacteriophages at Different pH Conditions
Briefly, 100 μL of phage solution (108 PFU/ mL) was suspended in 900 μL of SM buffer for studies on the stability of phages in acidic and alkaline conditions. The pH of SM buffer was adjusted with either 1 M HCl (Merck, Germany) or 1 M NaOH (Merck, Germany) to obtain solutions with pH ranging from 3 to 11 and incubated at 37°C. Samples were taken at 60 min and 24 h. The survival rate of each bacteriophage was determined at each pH value by the double-layer agar method (16).
Sensitivity of Isolated Phages to Ultraviolet Light
According to the method described by Allué-Guardia et al., 2012 with minor modifications, the phages were exposed to ultraviolet light (17). 2 mL of each phage suspension (108 PFU/ mL) was placed on a sterile petri dish under an ultraviolet lamp (λ= 254 nm) in a laminar hood (Caution, Iran) at a distance of 0.6 m from the radiation. Samples were taken at 0, 15, 30 min, and the phage survival rate was calculated.
The Effect of Osmotic Shock on Phages Survival
Evaluation of survival of phages during osmotic shock was done by incubation of viral particles in TM buffer (10 mM Tris–HCl (Merck, Germany) and 10 mM MgSO4 (Merck, Germany); pH=7.2) containing NaCl with the final concentration of 4.5 M. 0.1 mL of phage suspension (108 PFU/ mL) was mixed with 0.9 mL of TM buffer and incubated for 15 min at room temperature. Then, double-layer agar was prepared, and the number of phage plaques formed was counted. Phage suspension in TM buffer without NaCl was considered a control (18).
The Effect of Different Detergents on Phages Survival
To determine the antiviral effect of ionic detergents on phage particles, two different compounds were tested: anionic sodium dodecyl sulfate (SDS) (Merck, Germany), and cationic cetyltrimethylammonium bromide (CTAB) (Merck, Germany), according to the previously described method (18). Briefly, the phage suspension (108 PFU /mL) was mixed with 0.09% SDS solution and incubated at 45°C for 20 min. Also, phage suspension with the same titer was added to 0.1% CTAB solution and incubated at 22°C for 1 min. The formed plaques by the double-layer agar method were counted. As a control assay, the same process was performed exactly without adding ionic detergents.
The Effect of Different Organic Solvents on Phage Survival
The effect of four different organic solvents, including ethanol (Razi, Iran), acetone (Merck, Germany), chloroform, and dimethyl sulfoxide (DMSO) (Merck, Germany), was investigated on isolated bacteriophages. In summary, phage suspensions (108 PFU /mL) were added separately to various organic solvents, including 63% ethanol, 90% acetone, 75% chloroform, and 50% dimethyl sulfoxide. Mixtures containing ethanol and phage solutions as well as mixtures containing acetone and phage solutions were incubated for 1 h at 22°C. Chloroform mixtures, including each of the phage suspensions, were placed at 4°C for 1.5 h, and mixtures containing dimethyl sulfoxide and the phages were kept at 4°C for 10 min. Then, the plaques formed by the double-layer method were counted in each case. Phage suspension in SM buffer without organic solvents was used as a control (18).
Bacteriophage Host Range
To investigate the lytic activity of the isolated bacteriophages, their effects were also examined on the number of gram-negative and gram-positive bacteria additionally. A spot test was used to evaluate the susceptibility of bacteria to isolated phages. Pure culture of clinical isolates of P. aeruginosa, S. aureus, E. coli, and K. pneuminiae as well as standard strains of P. aeruginosa ATCC 27853, P. aeruginosa ATCC 9027, K. pneumoniae ATCC 1290, S. aureus ATCC 25923, and S. enterica ATCC 19430 and E. coli ATCC 8739 were prepared by the double-layer agar method. Of bacterial overnight cell culture, 100 μL was mixed with 3 mL of molten LB agar containing 0.7% agar and spread on an LB agar medium plate. The culture medium became hard after 15 min. Ten μL of phage stock was added as a spot on the plate surface of each used bacterial isolate. The plates were then incubated at 37°C overnight and examined for clear spots on bacterial cultures. Phage-free plates were used as negative control (19). To determine the host range of PPaMa3/19 and PPaMa4/19 phages, 22 P. aeruginosa bacteria (two standard strains, 20 clinical isolated from hospital specimens), 21 E. coli bacteria (one standard strain, 20 clinical isolated from hospital specimens), 21 K. pneuminiae bacteria (one standard strain, 20 clinical isolated from hospital specimens), 21 S. aureus bacteria (one standard strain, 20 clinical isolated from hospital specimens), and one standard strain of S. enterica were used.
Isolation of Phage
A lytic bacteriophage was successfully isolated against each of the reference strains of P. aeruginosa, producing clear plaques on the culture medium. These plaques have well-defined edges. Specific isolated phages against P. aeruginosa ATCC 9027 and P. aeruginosa ATCC 27853 were named PPaMa3/19 and PPaMa4/19, respectively.
Testing of the Optimal Multiplicity of Infection (MOI)
The results showed that the MOI rate of PPaMa4/19 was 0.1, and the highest titer obtained by this phage was 3.14 × 108 PFU/mL. The MOI of phage PPaMa3/19 was equal to 0.1, and the highest observed titer for this phage was estimated to be 4.21 × 108 PFU/mL. The MOI results for both phages are shown in Table 1.
Table 1. Optimal multiplicity of infection (MOI) of phages PPaMa3/19 and PPaMa4/19
| PPaMa3/19 (PFU/mL) | PPaMa4/19 (PFU/mL) | MOI | PFU | CFU/mL |
| 107 × 5.2 | 108 × 1.05 | 100 | 108 | 106 |
| 108 × 1.17 | 107 × 9.8 | 10 | 108 | 107 |
| 108 × 1.01 | 107 × 7.6 | 1 | 108 | 108 |
| 108 × 4.21 | 108 × 3.14 | 0.1 | 107 | 108 |
| 108 × 3.05 | 108 × 2.96 | 0.01 | 106 | 108 |
| 107 × 8.3 | 108 × 3.02 | 0.001 | 105 | 108 |
| 108 × 2.97 | 108 × 2.13 | 0.0001 | 104 | 108 |
Phage Adsorption Kinetics
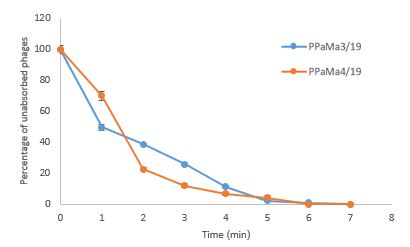
In this context, it was found that bacteriophage PPaMa4/19 is rapidly absorbed in P. aeruginosa cells. 77.28% of phages were absorbed by host cells after 2 min of mixing phage suspension with bacterial culture. After 6 min, 100% absorption rate was observed (Figure 1). Whereas 61.3% of PPaMa3/19 phages were absorbed by bacterial cells after 2 min of mixing phage suspension and bacterial culture, and after 7 min, 100% of the phages were absorbed by host cells (Figure 1).
One-Step Growth Curve
Both bacteriophages proliferated rapidly in the host cells with a short latent time. The host culture lysis was completed in 20-25 min. The latent time of PPaMa4/19 was five min, and its burst size was estimated to be 382 virions per infected cell (Figure 2). In the one-stage growth curve of PPaMa3/19 phage, a 6 min latent time was observed, and the burst size of this bacteriophage was 416 virions per infected cell (Figure 2).
Figure 1. Phage adsorption kinetics of PPaMa3/19 and PPaMa4/19 on host cells within 7 min.
Figure 2. One-stage growth curve of PPaMa3/19 and PPaMa4/19 phages over 25 min.
Characteristics of Plaques and Phages
In this study, the turbidity and diameter of plaques caused by bacteriophages were determined. Both phages formed clear plaques that are commonly seen in lytic phages. The diameter of plaques created by both phages was different. Figure 3 shows plaques created by both phages. The diameter of PPaMa3/19 plaques was about 1.5 mm, and the diameter of plaques produced by PPaMa4/19 phage was about 2.5 mm.
Figure 3. (A) Plaques formed by bacteriophage PPaMa4/19 and (B) plaques formed by phage PPaMa3/19.
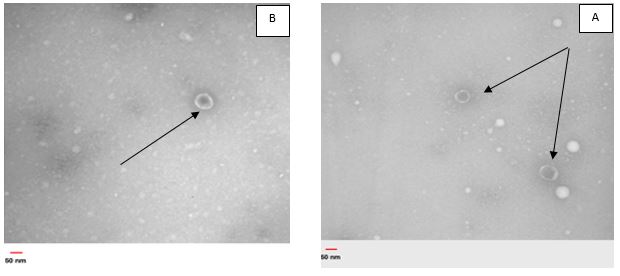
The morphological characteristics of both phages were tested by electron microscopy. Both had isometric heads and no tails. The average internal diameter of the PPaMa4/19 phage was 60 ± 2 nm, and the external diameter was about 70 ± 2 nm (Figure 4. a). The internal diameter of PPaMa3/19 phage was 65 ± 2 nm, and the external diameter was 78 ± 2 nm, as shown in (Figure 4. b). Due to the shape characteristics and phage size, both bacteriophages were most similar to the Cystoviridae family.
Figure 4. Photos taken by electron microscopy. (A) Specific phages isolated against Pseudomonas aeruginosa PPaMa4/19. (B) Specific phages isolated against Pseudomonas aeruginosa PPaMa3/19. Both photos have a magnification of 50 nm.
Stability of Bacteriophage Against Temperature and pH
Both isolated phages were tested over a 24 h period against a range of pH= 3-11 to determine their stability under acidic and alkaline conditions (Figure 5). Both phages showed good stability at pH= 5-9, and the highest resistance was observed at pH= 7. Phage PPaMa3/19 showed more stability than phage PPaMa4/19 at acidic pH, and PPaMa4/19 phage was more stable than PPaMa4/19 phage at alkaline pH. After 24 h, no stability of PPaMa4/19 was observed at pH=3. The stability of PPaMa4/19 in alkaline conditions was higher than PPaMa3/19 in overtime. The results related to the stability of phages to different temperatures have been measured and are shown in Figure 6. The highest stability of both phages was observed at 4°C. With increasing temperature and time, the titers of both phages decreased compared to the primary titers. The greatest decrease in the stability of both phages was at 60°C. At temperatures of -20, 22, and 37°C, a slight decrease in phage survival was observed.
Figure 5. Survival rate of PPaMa3/19 and PPaMa4/19 bacteriophages in acidic and alkaline condition.
Figure 6. The survival rate of PPaMa3/19 and PPaMa4/19 bacteriophages in temperatures of-20, 4, 22, 37, 40 and 60°C.
Resistance to Ultraviolet Light
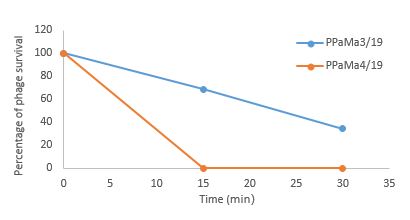
Both PPaMa3/19 and PPaMa4/19 bacteriophages were exposed to ultraviolet radiation for 15 and 30 min. The results are shown in (Figure 7). PPaMa4/19 was more sensitive to UV than PPaMa3/19. The survival rate of PPaMa4/19 phage was zero after 15 min. The PPaMa3/19 titer also showed a significant decrease after 15 and 30 min of UV exposure. Eventually, in the first 15 min the survival rate of phage was 68.7%, and during the second 15 min it was decreased to 34.53%.
Figure 7. Resistance of PPaMa3/19 and PPaMa4/19 phages against ultraviolet rays.
The Effect of Osmotic Shock on Phage Survival
Both phages showed a high survival rate against these conditions. Phage PPaMa3/19 above 99%, and phage PPaMa3/19 showed above 73% resistance to saline solution. The corresponding results are shown in Table 2.
Effects of Detergents and Organic Solvents on Phage Survival
In general, detergents (sodium dodecyl sulfate and cetyltrimethylammonium bromide) and organic solvents (ethanol, acetone, chloroform, and dimethyl sulfoxide) caused a decrease in titers of both phages. The survival rate of PPaMa3/19 in the presence of chloroform was 50.66 %, and PPaMa4/19 was 40.2%. Responses of both phages to SDS, CTAB, and DMSO were quite different. However, PPaMa3/19 was more resistant to the organic solvents and detergents used in this study than PPaMa4/19, indicating the high resistance of this phage to various environmental conditions. In this study, ethanol and acetone were the main anti phage agents because both phages were more or less sensitive to this solvent. The results of this section are shown in Table 2.
Table 2. Percentage of stability of PPaMa3/19 and PPaMa4/19 phages against osmotic shock, detergents, and various organic solvents used in this study.
| Phage survival rate under study conditions (percentage of phage survival) | |||||||
| Phages | CTAB (0.1) |
SDS (0.09%) |
DMSO (50%) |
Acetone (90%) |
Chloroform (75%) |
Ethanol (63%) |
Osmotic shock |
| PPaMa3/19 | 46.6% | 49% | 87.7% | 20.8% | 50.66% | 46.6% | 99% |
| PPaMa4/19 | 0 | 0 | 4.8% | 0 | 40.2% | 0 | 73% |
Bacteriophage Host Range
The PPaMa3/19 and its specific host lysed the standard P. aeruginosa 27853 ATCC strain and a clinical isolate of P. aeruginosa among 86 bacteria from different genera. This phage had a very limited host range, which indicated the high specificity of this bacteriophage. In return, phage PPaMa4/19 showed lytic ability against 23 of 86 isolates, including 2 standard strains of P. aeruginosa 27853 ATCC, P. aeruginosa 9027 ATCC, 9 clinical isolates of P. aeruginosa, 8 clinical isolates of E. coli and 4 isolates of S. aureus.
Due to the importance of combating antibiotic-resistant nosocomial infections caused by P. aeruginosa, the use of effective methods in this field is one of the priorities of clinical research; therefore, finding effective alternative methods to control these bacteria is significant. In the meantime, the use of the antibacterial potential of phages has been considered due to their cheap isolation and their unique characteristics against bacteria. In this context, the high specificity between the phage and the host indicates the need to isolate new phages that can remove large numbers of multidrug-resistant bacterial strains of P. aeruginosa, and no potential phage has been identified to eliminate/ kill all P. aeruginosa species as of yet.
In the present study, isolation of lytic phages against two reference strains of P. aeruginosa as hosts and assessment of the properties of these phages. Also, measurement of their antibacterial potential against clinical P. aeruginosa infections as well as some other bacteria isolated from clinical infections (Gram-positive and Gram-negative bacteria) was evaluated. After successful isolation of two specific bacteriophages and purification, a one-step growth curve of both phages was plotted. The one-step growth curve is the curve resulting from changes in phage amplification in the host (20). Through it, valuable information such as the burst size and the latent time of the bacteriophages can be obtained. The burst size is equal to the number of viral particles released during the lysis of the host cells, which is an important parameter in viral ecology.
In general, lytic phages have a shorter latent time and a smaller burst size than temperate phages (21). One of the remarkable properties of the two bacteriophages isolated in this study was their rapid growth in P. aeruginosa cells under standard laboratory conditions with a very short latent time of 4 to 5 min. The very rapid proliferation of lytic phages with a relatively large burst size indicates the high potency of these bacteriophages in killing bacteria. In 2021, Yang et al. isolated phage KPP10 from river water and used it against pneumonia caused by P. aeruginosa (22).
In another study conducted by Topka et al. in 2019, phage vB_EcoS-95 had a short latent time of 4 min with a burst size of 115 phages per cell, and after 2 min of mixing the phage suspension with the host culture, about 50% of the phages were absorbed the bacteria (12). According to the one-stage growth curve of AZ1 phage isolated against P. aeruginosa by Jamal Muhsin et al. in 2017, a long latent time of 33 min and a burst size of over 326 phages per cell were reported for this phage (5). In 2018, Forti et al. used a phage cocktail to treat acute P. aeruginosa infections. All isolated phages belonged to the order Caudovirales. One-step growth experiments of the respective phages showed that the PYO2 and DEV phages had the same latent time (20 min) and burst sizes of 200 and 100 PFU/ mL, respectively. While E215 and E217 had a long latent time (30 to 40 min) and a burst size of more than 200 PFU/ mL (23). The isolated phages in the present study produced plaques of different sizes. The results of electron microscopy showed that the phages were polyhedral without tail, and both phages belonged to the Cystoviridae family. In this regard, Shukri et al., 2016, isolated two bacteriophages belonging to the Cystoviridae and Leviviridae families against antibiotic-resistant clinical strains of P. aeruginosa from Isfahan hospital wastewater (24). Phage phi6 is the first phage belonging to the Cystoviridae family isolated by Yang et al. in 2016 against P. aeruginosa PAO38 (25). In 2019, Jeon and Yong investigated the potential of two bacteriophages, Bφ-R656 and Bφ-R1836, to control the clinical strains of P. aeruginosa XDR-PA. Both isolated phages belonged to the Syphoviridae family and had a wide host range against different strains of P. aeruginosa (26).
The practical use of phages in different contexts requires their stability under different tension conditions. Acidic or alkaline environments, high or low temperatures, and various chemicals can affect the function or survival of phages. Therefore, determining the stability of bacteriophages in different conditions to select an effective phage is of particular importance for using their antibacterial potential in different fields. In the present study, PPaMa3/19 and PPaMa4/19 phages had high stability between pH 5 to 9, and the highest stability for these bacteriophages was observed at pH= 7. Phage AZ1 was stable at pH 3 to 11, had the highest stability at pH= 7, and was inactivated at pH 1 and 2 (5).
In a study conducted by Litt et al., in 2017, of the seven isolated phages, three phages were stable at pH= 1 and all seven phages maintained their stability at pH= 11-2 after 24 h (15). Adnan et al., in 2019, isolated MA-1 against P. aeruginosa with a moderate host range from sewage. This phage of the Myoviridae family was stable against different temperatures and pH values so that it had a suitable lytic activity at a temperature of 37 to 65°C and was completely inactive at temperatures of 80°C and above. Phage MA-1 was stable at pH 5 to 9 and decreased its activity at pH=3 and 11 (27).
In another study, Feng et al., in 2003, examined the stability of 2 phages, MS2 and Qβ. Phage MS2 showed a better survival rate in acidic conditions and phage Qβ in alkaline conditions. The titers of both bacteriophages were significantly reduced at pH below 6 and above 8 (28). Strack et al. showed a linear relationship between mutation rate and incubation of phages at low pH in this context. This relationship suggested that phages could mutate to survive in an acidic environment (29). Checking the stability of phages at different temperatures is essential to ensure their survival during storage and transport. In the present study, a decrease in phage activity was observed at 60°C, and between -20 and 60°C; the most suitable temperature to store both isolated phages was 4°C. In 2017, Pushpinder et al. did not observe a significant reduction in phage activity at 60°C for 60 min, and five of seven isolated phages in the study survived at 70°C for 10 min. Also, all phages were active at 4, -20, and -80°C for 90 days (15). In a study by Wang et al. in 2016, the SLPW maintained its stability up to a maximum of 45°C, and higher temperatures led to the gradual inactivation of this phage. At 65°C, the phage was completely inactivated (11).
In 2019, Guo et al. investigated the effect of different temperatures and pH values on two lytic phages isolated against P. aeruginosa. Phage S1 was more stable to pH changes than phage S2. Both phages exhibited high lytic activity between pH 4 to 10 for one hour. Still, they behaved quite differently to temperature changes so that the stability of phage S1 gradually decreased by increasing temperature up to 60°C and completely disappeared at 80°C. A slight decrease was observed in the stability of phage S2 by increasing temperature up to 60°C, but completely disappeared at 70°C or more. Different responses of two phages to similar temperatures can be due to existing distinct proteins or their capsid structures (30). According to research, different bacteriophages show different behaviors against different temperature treatments (24), which can explain the differences in the results of the studies. Ultraviolet radiation is another environmental factor affecting microorganisms that also reduce the infectivity of phages. As a result, it reduces their potential use as biocontrol agents (31). According to a study by Allué-Guardia et al. in 2012, phages were sensitive to UV exposure, and this radiation has reduced the phage titer between log 3-4. It has caused complete inactivation of phages (17) even after 30 min. The results obtained in the present study were consistent with the results of previous studies. During the present study, the effect of osmotic pressure on phages was also investigated by diluting the phage suspension with a high salt concentration. Both phages showed a high survival rate (73-100%). In a study by Jurczak et al., in 2016, out of 83 isolated phages, 62 phages showed high survival (70-100%) to osmotic shock, and 21 phages were sensitive to it (18).
The organic solvents and detergents used in this study caused a decrease in phage titer, which had the highest anti-phage activity of ethanol and acetone. Both phages showed the highest sensitivity to these organic solvents. In a 2016 study, acetone had the most anti-phage activity on phages, and sodium dodecyl sulfate, cetyltrimethylammonium bromide, and dimethyl sulfoxide caused a significant decrease in the titer of most of the phages in this study. Also, a limited number of phages showed a high survival rate against organic solvents and detergents. One of these phages was vB_Eco1M, which survived even at high temperatures (18). Bacteriophages are more tolerant to alcohol than bacteria (32). Due to the use of chloroform in different stages of isolation, it was necessary to investigate the sensitivity of phages to chloroform, and it was found that both isolated phages were sensitive to chloroform. Studies have shown that Cystoviridae phages are sensitive to chloroform because of their membrane. Based on our results, PPaMa4/19 showed a wider host range than PPaMa3/19. This way, PPaMa3/19 was only able to lysis 2 reference strains and 1 clinical isolate of P. aeruginosa and had no effect on the other used clinical isolates while the PPaMa4/19 had a lysis effect on both reference strains and 9 isolates of P. aeruginosa and a number of E. coli and S. aureus isolates, but none of the bacteriophages had lytic effect on the isolates of K. pneumoniae and S. enterica. Muhsin Jamal et al., examined the effect of phage AZ1 on 34 different strains. This phage was able to lysis only 6 bacterial strains, indicating a limited host range of this phage (5).
Knezevic et al., in 2009, also examined host range of 19 phages isolated against different P. aeruginosa on 32 strains of P. aeruginosa and 5 different strains of Enterobacteriaceae family (S. enterica ATTC 13076, K. pneumoniae ATCC 13883, Proteus mirabilis Pm-9, E. coli ATCC 25922 and ATTC 8739). None of the isolated phages was able to lysis the five mentioned strains (33). Melo et al., tested the effect of BrSP1 phage on 37 strains of P. aeruginosa. This phage showed the ability to lysis 19 strains (51.4%) (34). Host range of vB_PaeP_PPA-ABTNL isolated against antibiotic resistant P. aeruginosa on 15 isolates of P. aeruginosa, P. aeruginosa ATCC 27853, E. coli O157: H7 ATCC35150, K. pneumoniae, P. mirabilis, S. aureus, S. enterica, and Listeria monocytogenes were analyzed by Cao et al., in 2015. The phage was able to remove only 14 P. aeruginosa isolates and had no effect on the standard P. aeruginosa ATCC27853 strain (35).
In a study by Sepúlveda-Robles et al., in 2012, to study phage diversity in central Mexico, 68 isolates were isolated against seven strains of P. aeruginosa and investigated the lethal effect of each phage on 142 strains of P. aeruginosa. Lysis patterns were recorded for 112 strains susceptible to phage infection. Then, phages and bacteria were analyzed with similarity host range by using the software program (36). Yang et al. in 2020, prepared a phage cocktail of 5 phages. It's capable of lysis a wide range of clinical strains of P. aeruginosa. This study also showed that using the phage cocktail can eliminate the problem of phage resistance caused by the strains and increase the efficiency of individual phages in the phage cocktail (37). The use of bacteriophages can play an effective role in combating bacteria due to their abundance, cheapness and accessibility.
Bacteriophages have less destructive effects on the natural flora of the body due to their specific function. Unlike antibiotics, this high specificity prevents the effect of phage lysis on a wider range of bacterial pathogens. In various studies, the antibacterial power of phages against bacteria from other bacterial genera and their specific host has been identified. This potential of bacteriophages can also fight infections caused by bacterial mixtures. In addition, for therapeutic use of the antibacterial potential of isolated phages, various factors such as host range, phage durability in different environmental conditions, and their other biological properties should be evaluated and the results of the properties of one phage can not necessarily be generalized to other phages. In order to get more accurate identification of phages, determining their nucleotide sequences is essential and tangible to investigate the absence of pathogenic genes. Unfortunately, these assays were not done in present study due to the lack of necessary facilities, budget, and the impossibility of doing it in our country.
Based on the present study's findings, two bacteriophages isolated against standard strains of P. aeruginosa 27853 ATCC and P. aeruginosa 9027 ATCC as hosts had significant and strong lethal effects on their hosts. The high absorption rate, short latent time, and high burst size of both phages indicate this fact. The isolated phages belonged to the family Cystoviridae, and showed stability in the temperature range between -20 to 37°C or the pH range 5 to 9. These characterizations indicate the ability to use the phages for phage therapy in terms of appropriate antibacterial potential and suitable stability in environmental conditions. The organic solvents and ionic detergents used in this study decreased the titers of both phages; the highest anti-phage activity belonged to ethanol and acetone, which should be considered in the use of these phages.
Despite the above mentioned, investigating the host range of bacteriophages against bacteria is another important issue that should be considered before using these phages in phage therapy. In this study, PPaMa4/19, unlike PPaMa3/19, had antibacterial activity against E. coli (40%) and S. aureus (20%). The other phage had a narrow host range and only had an antibacterial effect against its host and one of P. aeruginosa clinical isolates. So, in general, it can be said that each of the two phages PPaMa3/19 and PPaMa4/19, with suitable antibacterial properties, has adequate potential for therapeutic use. However, further studies such as sequencing of the entire phage genome, molecular and bioinformatic analysis of phage sequences, a study of phage protein content, clinical experiments, and evaluation of phage stability in various other environmental conditions for the applied use of these bacteriophages are inevitable.
Not applicable.
None.
Conflicts of Interest
The authors declared no conflict of interest.
Received: 2021/04/27 | Accepted: 2022/02/9 | ePublished: 2022/05/25
| Rights and permissions | |
 |
This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License. |